Machine Learning is disrupting the world of medicine and healthcare, allowing professionals to diagnose patients better and faster than before. However, any Healthcare AI ML model training needs high-quality annotated medical images in large quantities. This is where medical data labeling becomes essential. By leveraging medical imaging services and medical imaging tools specifically designed for annotation, healthcare AI teams can ensure precise data preparation. This blog delves into the fundamentals of medical image annotation, covering everything from dataset preparation to the technical considerations unique to healthcare.
For professionals and teams in AI-driven healthcare, understanding the intricacies of labeling medical data is crucial to advancing diagnostic accuracy and patient outcomes. Let’s dive in to learn more.
What is Medical Image Annotation?
Medical image annotation, also known as medical image labeling is the process of annotating medical data, be it imaging data such as CT scans, X-rays, MRIs, ultrasounds, and retina fundus shots so that healthcare professionals and machine learning algorithms can accurately interpret and diagnose medical conditions, track disease progression, and make informed treatment decisions.
The healthcare industry also requires other types of data to be labeled, such as document data like medical records in PDF or PNG/JPG formats. Medical data labeling can include audio labeling, such as patient conversations or cough sounds. This blog will focus on medical imaging.
AI teams use labeled data to train their ML models, which, once trained, can then automatically detect objects, lesions, tumors, and other abnormalities.
The Importance of Medical Data Annotation
In addition to imaging data, the healthcare industry requires other types of data to be labeled, such as document data (medical records in PDF or PNG/JPG formats) and audio labeling (patient conversations or cough sounds). While this blog focuses on medical imaging, understanding the broader context of medical data annotation is essential, as it encompasses a variety of tasks that support AI in healthcare.
Medical Data Annotation vs. Medical Image Annotation
Medical data annotation is a broad term that covers the labeling of various types of medical data, including imaging data (like CT scans, MRIs, and X-rays), document data (such as patient records in PDF or image formats), and even audio data (like patient conversations or cough sounds). Each type plays a critical role in helping AI systems learn to recognize patterns and support healthcare decision-making.
In contrast, medical image annotation specifically refers to the labeling of imaging data for machine-learning purposes. While this blog focuses on medical image annotation, we recognize that medical data annotation is a wider concept that iMerit supports with a platform designed to handle diverse challenges and requirements.
Key Considerations for Effective Medical Image Labeling
Implementing a labeling strategy that covers diverse imaging formats and real-world application requirements is vital. Medical data often exists in varied formats, such as DICOM for radiology or TIFF for microscopy, which demand specialized medical imaging tools for accurate annotation. Ango Hub platform is compatible with medical imaging services, supporting a wide range of formats to streamline the labeling process and ensure high accuracy.
On the other hand, it is crucial to get the images ready for labeling. In order to train a machine learning model that can produce dependable results, it is crucial to provide it with a substantial amount of high-quality labeled data. Frequently, obtaining this data, even in its unlabeled form, can be challenging. Additionally, when you have access to the data, there are a couple of considerations to keep in mind.
1. Building a Strong Dataset for Machine Learning Model Training
The success of a medical AI model often depends on both the size and variety of its training dataset. Incorporating images from multiple sources ensures that the model can generalize effectively, reducing the likelihood of bias and enhancing its performance in real-world applications. By blending data from various sources, it is possible to create a model capable of providing accurate predictions across a wider range of patients.
2. Variety of Datasets
Ensuring data diversity is crucial; it should not originate solely from one source or exhibit uniformity in appearance. The goal is to make the model as robust as possible to handle a wide range of real-world scenarios. If the model was trained on a subset of data that closely resembles each other, it might struggle when presented with diverse data. In essence, incorporate data from various sources, stages, institutions, or locations to enhance the model’s adaptability to different situations.
3. Data Quality and Annotation Standards – It Matters
Maintaining high standards for data labeling can dramatically improve the model’s efficacy. Enforcing quality checks and review processes ensures that each annotation meets stringent accuracy requirements, especially in healthcare, where precision is paramount. This practice of double-checking annotations often proves indispensable for medical imaging, where nuanced details like lesion boundaries are crucial.
4. The Dataset Vetting Process
We recommend splitting your dataset into training, validation, and testing, where the training dataset should comprise about 80% of your data. First, train your model with the training set and then evaluate the results on a small validation set. Look at the results that come out of the validation set. Are they to your satisfaction? Chances are, some adjustments will be needed. Make the necessary tweaks, retrain, and validate again. Repeat this process until the validation results meet your expectations. Once you are happy with the validation results, test your results against the test dataset. It will be your final model benchmark.
5. Balancing Quantity and Quality for Model Optimization
While a larger dataset is often preferred, high-quality datasets are crucial for accurate annotations. Smaller datasets of high-quality, accurate annotations can outperform larger but less precise datasets. Expanding datasets only when maintaining quality is feasible ensures that resources are utilized effectively and improves model robustness without unnecessary complexity.
6. Size of your Dataset
Recent developments in ML have shown that quality is as important as quantity in training models. It means that a smaller but high-quality set will usually perform equally or even better than a larger set of lower quality. That said, if you have the option to enlarge your dataset, we highly recommend doing so, as model results will improve significantly.
7. Format of your Dataset
The two most common medical imaging formats around are DICOM and TIFF. DICOM, especially, is the industry standard for radiologists. These files can optionally contain multiple images, slices, and metadata regarding the patient and the image itself.
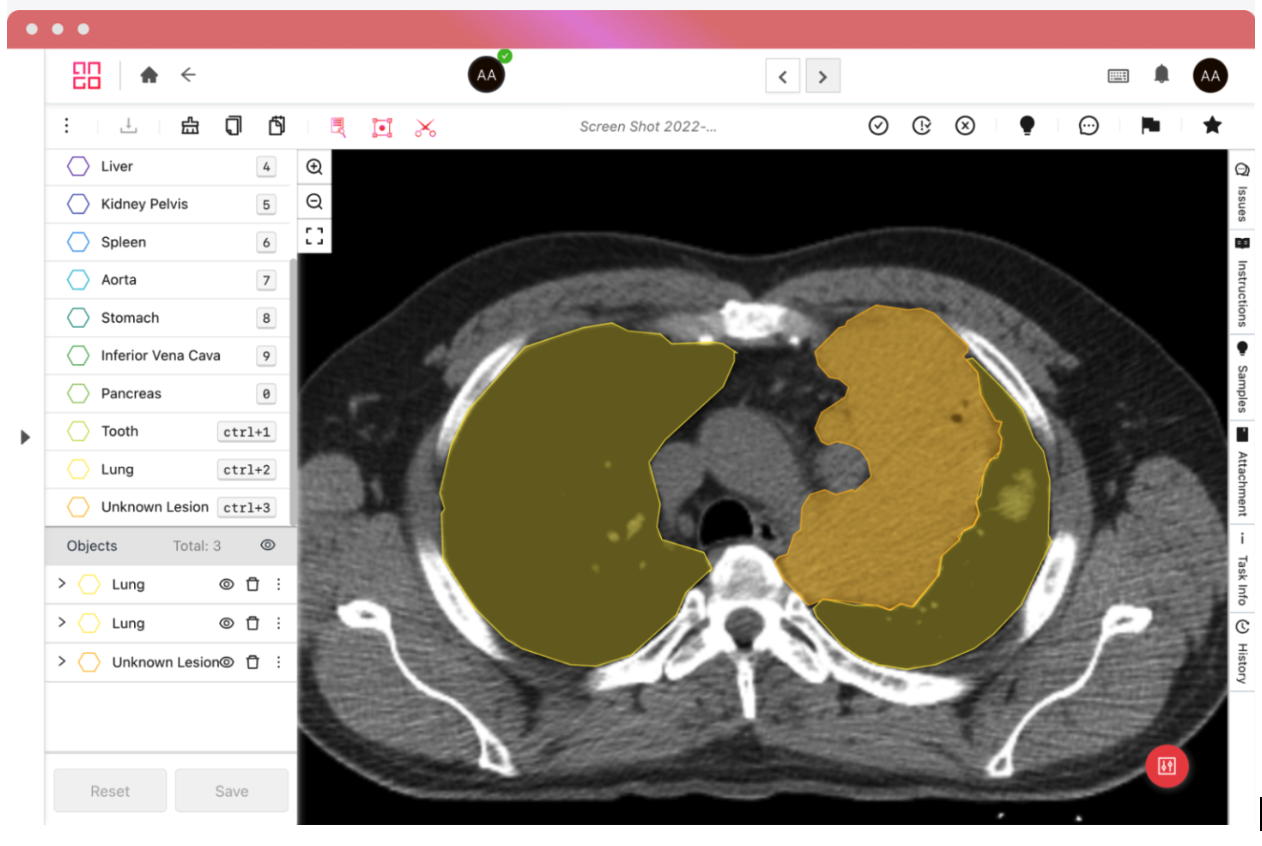
Good medical data annotation platforms will support both these formats, and the iMerit Radiology Editor, powered by the Ango Hub, can automatically remove identifying information from both metadata and the image itself on upload.
What Makes iMerit’s Medical Data Annotation Different From Others?
Labeling images for healthcare is an altogether different endeavor compared to regular image annotation.
Here are some things that are different:
Data Availability
While regular images are often freely available or behind a standard NDA, medical imaging is usually protected by strict data processing agreements. It is mainly to protect the privacy of the patient. Obtaining medical imaging data is usually a longer process than other data types.
Technical Differences
Regular images only have one layer, are of small size, and have a low bit depth. Medical images often have multiple layers (slices), are huge, and have a higher bit depth. Further, the labeler profiles for both will be different, where the annotation of medical images demands expertise from specialized healthcare professionals. These experts are used to certain UI and UX paradigms. Therefore, when choosing a data labeling platform, it is critical to note whether medical professionals can easily use its keyboard controls and UI.
Domain Experts Matter
Medical image annotation isn’t just about labeling, it’s about labeling correctly. iMerit’s Domain Experts and Scholars Program provides trained professionals, including radiologists and clinical specialists, to ensure annotations meet clinical-grade standards. This expert-in-the-loop approach dramatically improves output quality and model trustworthiness.
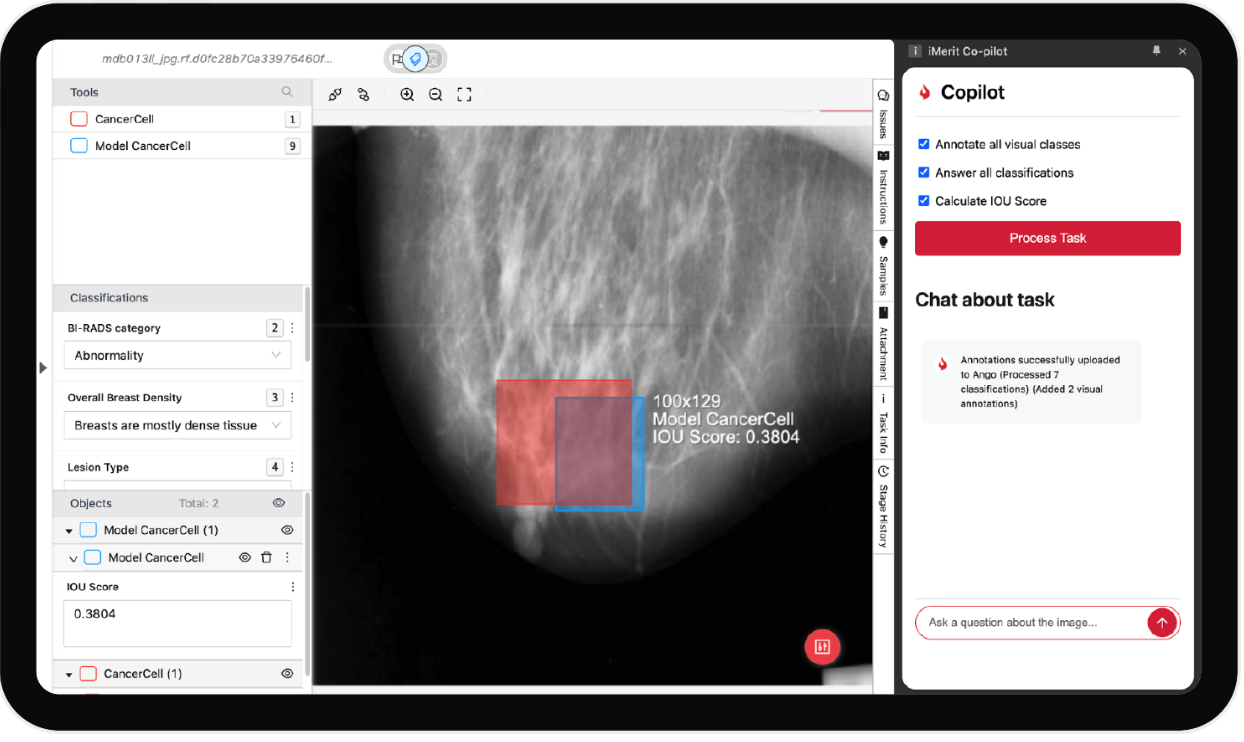
Radiology Annotation Co-Pilot
The Radiology Annotation Co-Pilot enhances productivity by assisting annotators in real-time. It automates repetitive tasks, suggests annotations based on prior labels, and learns from expert corrections to reduce turnaround time and human error, especially in complex labeling like segmentation or classification.
Driving ML Ops Efficiencies Through Workflow Designer
Workflow automation is crucial to scale annotation operations. Ango Hub’s Workflow Designer allows teams to create, manage, and iterate on structured labeling processes from queue assignments and QA routing to domain expert escalation. This supports faster annotation cycles and enhances ML model training timelines.
Reports and Analytics
Healthcare AI teams require visibility into annotation performance, workforce productivity, and data throughput. Ango Hub includes built-in reporting and analytics dashboards that track progress, quality scores, labeling velocity, and audit results, empowering continuous improvement and transparency.
Data Security
Healthcare data privacy is non-negotiable. iMerit complies with HIPAA and other regulatory frameworks. Ango Hub offers:
- Automated PHI removal
- Role-based access controls
- Secure on-prem, cloud, or hybrid deployment
- Full audit trails
This makes it ideal for handling sensitive medical datasets safely.
Flexible Deployment: On-Shore, Off-Shore, Hybrid Teams
Whether you need an on-shore annotation team for compliance reasons or a hybrid setup to balance cost and speed, iMerit offers flexible workforce models. Teams are trained in medical protocols and can be deployed globally based on your data security and project needs.
On-Prem vs Cloud
Some healthcare organizations require on-premises deployment due to internal IT policies or sensitive data handling. Ango Hub supports on-prem, cloud, and hybrid models, ensuring that AI development doesn’t stall due to infrastructure limitations.
Advanced Features That Drive Quality
The combination of expert input, co-pilot automation, real-time QA, and PHI compliance makes iMerit’s platform ideal for medical annotation. The features below, all contribute to faster, more accurate medical image annotation that is model-ready.
- Smart pre-labeling
- Instant feedback loops
- Multi-modality support
- Multi-slice navigation
- AI-powered annotation suggestions
Picking the Medical Data Annotation Tool for You
DICOM viewers with annotation capabilities abound in the market. One notable open-source option, for example, is 3D Slicer. DICOM viewing tools, however, are not optimized for ML model training. Sometimes, it is impossible to use the labels from these viewers in machine learning due to a lack of instance IDs and structured export formats. You must use a professional medical imaging labeling tool to train and develop a neural network.
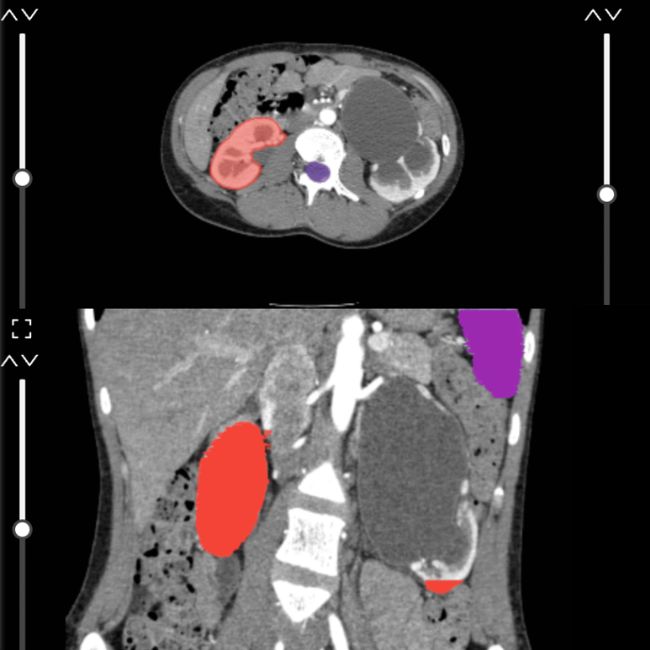
Answer below for the image annotation solution you use or are choosing:
- Does the solution support medical formats such as DICOM and TIFF?
- Does it support the labeling features you are looking for?
- Is the UX easy to use and suitable for medical use?
- Is the export format easy to use in ML model training?
- Does the tool provider have a medical data labeling service to enhance your workforce?
Conclusion
High-quality medical data annotation is essential to the success of machine learning applications in healthcare. From dataset diversity and quality to specialized tools like Ango Hub, each aspect of the annotation process supports building accurate, reliable AI models that enhance diagnostics and treatment. iMerit’s robust medical imaging services and medical imaging tools ensure that annotated data is optimized for AI applications, making it a key resource for any healthcare AI project.
As healthcare AI continues to advance, understanding the intricacies of data labeling will empower professionals to drive more precise outcomes across the field. If you’re ready to elevate your medical image annotation project, sign up for iMerit’s Ango Hub and start labeling your medical images with confidence.
